 Software applying machine learning has become an essential tool to automate analysis, but these methods require annotated examples to learn from. High-content, imaging-based screens now routinely generate data on a scale that precludes manual verification and interrogation. ilastik is an user-friendly interactive tool for machine-learning-based image segmentation, object classification, counting and tracking. We describe all ilastik workflows in detail, including three case studies and a discussion on the expected performance. Once the classifiers are trained, ilastik workflows can be applied to new data from the command line without further user interaction. Its computational back end runs operations on-demand wherever possible, allowing for interactive prediction on data larger than RAM. ilastik can process data in up to five dimensions (3D, time and number of channels). Users adapt the workflows to the problem at hand by interactively providing sparse training annotations for a nonlinear classifier. It contains pre-defined workflows for image segmentation, object classification, counting and tracking. We present ilastik, an easy-to-use interactive tool that brings machine-learning-based (bio)image analysis to end users without substantial computational expertise. 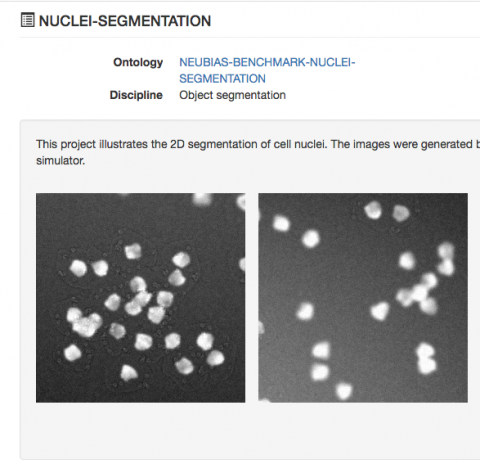
We hope to expand awareness of the powerful deep-learning tools available to biologists for image analysis. In addition to surveying these tools, we overview several challenges that remain in the field. These tools take many different forms, such as web apps, plug-ins for existing imaging analysis software, and preconfigured interactive notebooks and pipelines. Here, we survey several new open-source software tools that aim to make deep-learning–based image segmentation accessible to biologists with limited computational experience. Until the past few years, however, most deep-learning workflows required significant computational expertise to be applied. By learning from large amounts of annotated data, deep learning can accomplish several previously intractable bioimage analysis tasks. However, extracting this complex information can be challenging, especially when biological structures are closely packed, distinguished by texture rather than intensity, and/or low intensity relative to the background. Microscopy images are rich in information about the dynamic relationships among biological structures. Supplementary Data 1: Benchmarking performance of classifiers in CPA 2.0 versus CPA 1.0 Supplementary Information: Supplementary Text 1: Manual to CellProfiler Analyst updated versions are available at /CPA We implemented an automatic build process that supports nightly updates and regular release cycles for the software. It is available as a packaged application for Mac OS X and Microsoft Windows and can be compiledįor Linux. CellProfiler AnalystĢ.0, completely rewritten in Python, builds on these features and adds enhanced supervised machine learning capabilities (Classifier),Īs well as visualization tools to overview an experiment (Plate Viewer and Image Gallery).Īvailability: CellProfiler Analyst 2.0 is free and open source, available at and from GitHub () under the BSD license. We hope these changes will make CellProfiler an even better tool for current users and will provide new users better ways to get started doing quantitative image analysis.ĬellProfiler Analyst allows the exploration and visualization of image-based data, together with the classification of complexīiological phenotypes, via an interactive user interface designed for biologists and data scientists. We’ve also added more explanations to CellProfiler’s settings to help new users get started. We’ve also made changes to CellProfiler’s underlying code to make it faster to run and easier to install, and we’ve added the ability to process images in the cloud and using neural networks (deep learning). In this release, we’ve added the capability to find and measure objects in three-dimensional (3D) images. Pipelines are easy to save, reuse, and share, helping improve scientific reproducibility. 
Researchers can download an online example workflow (that is, a “pipeline”) or create their own from scratch. The third major release of our free open-source software CellProfiler is designed to help biologists working with images, whether a few or thousands. Thus, many biologists find they need software to analyze images easily and accurately. 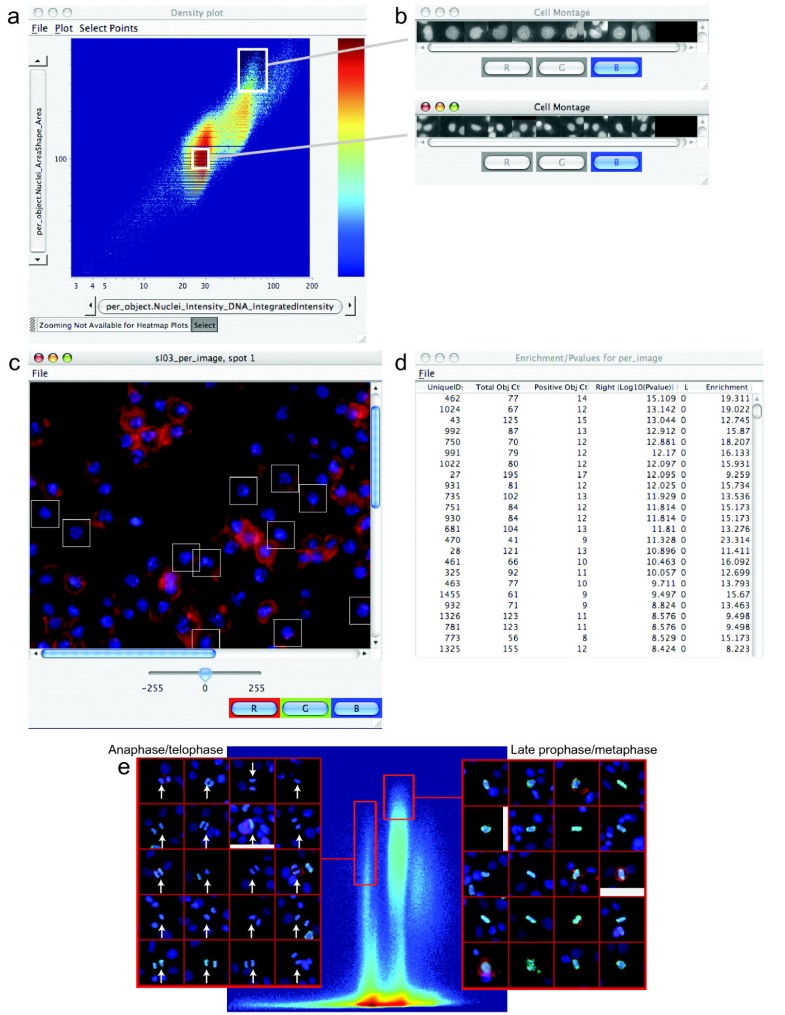
Looking at the resulting images by eye would be extremely tedious, not to mention subjective. The “big-data revolution” has struck biology: it is now common for robots to prepare cell samples and take thousands of microscopy images. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
